Source: Frontiers in Microbiology (2025) — DOI: 10.3389/fmicb.2025.1662087
Authors: Chao Wei, Yaxiang Wang & Zhe Chen
Brief Abstract:
This large-scale study profiled prophages, viral elements integrated into bacterial or archaeal genomes, across the porcine gut microbiome. By screening over 7,500 gut-derived microbial genomes, the authors uncovered more than 10,000 prophage genomes and revealed how these viral entities shape host defense, metabolism, and virulence. The findings highlight the extensive diversity and ecological importance of prophages as silent but influential players in the pig gut ecosystem.
What the Researchers Did (Methods & Materials):
- Dataset: 7,524 high-quality bacterial and archaeal genomes isolated from pig intestines and feces.
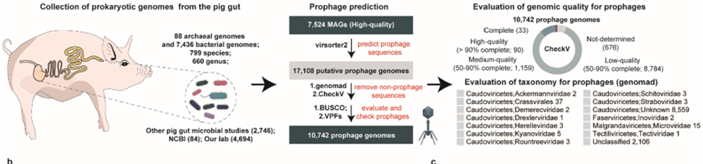
- Identification tools: VirSorter2, CheckV, and geNomad for prophage prediction and validation.
- Functional analysis: CRISPR spacer mapping to define host range; DefenseFinder to identify antiviral systems; VIBRANT and DRAM-v to detect auxiliary metabolic genes (AMGs); and CARD/VFDB databases for antibiotic resistance (ARGs) and virulence factor (VFG) genes.
- Comparative clustering: Genomic comparisons with global porcine and human gut virome databases to assess novelty and diversity.

Key Findings:
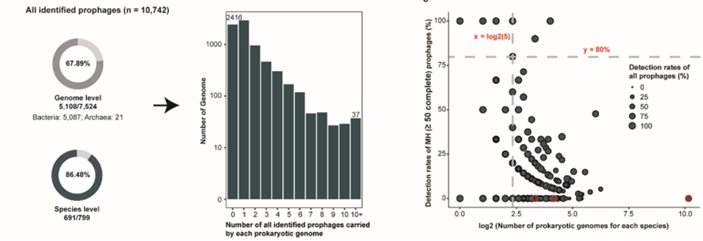
- 10,742 prophages identified across pig gut microbes; ~68 % of genomes carried at least one prophage.
- 1.7 % exhibited broad host-range infectivity, including cross-phylum potential.
- 5 % encoded defense systems enhancing host resistance via CRISPR-Cas or restriction–modification genes.
- AMGs: Detected in 1.1 % of prophages, notably cobA, cobS, and cobT genes promoting vitamin B₁₂ biosynthesis.
- ARGs: 208 genes found in 120 prophages, mainly multidrug, aminocoumarin, and nitroimidazole resistance.
- VFGs: 1,111 genes detected across 378 prophages – dominated by capsule, flagella, and LPS-associated functions.
- Integrase and tail-fiber proteins identified as major determinants of host specificity.
- Novelty: Over 88 % of prophages were previously uncharacterized, significantly expanding known diversity within Caudoviricetes and crAss-like phages.
Why It Matters to us:
- Prophages are not passive genomic remnants – they actively influence the structure and function of the gut microbiome.
- Their ability to transfer metabolic, antibiotic resistance, and virulence genes makes them crucial to understanding:
- The evolution of gut microbial communities in pigs.
- Mechanisms of antibiotic resistance spread.
- Potential for leveraging prophages in precision microbiome management and phage-based feed additives.
Headline Takeaway:
Hidden viral passengers in the pig gut drive bacterial defense, vitamin B₁₂ synthesis, and antibiotic resistance – reshaping our understanding of microbial evolution and resilience.
